Kazusa Clone (Original Type)
Gene/Protein Characteristic Table for
ORK01452
|
Order Kazusa clone(s) from : |
| Accession No | |
|---|---|
| Product ID | ORK01452 |
| Clone name | bm01873 |
| Vector information | |
| Symbol | STIP1 |
| cDNA sequence | DNA sequence (1810 bp) Predicted protein sequence (543 aa) |
|
HaloTag ORF Clone |
FHC01452

|
| Flexi ORF Clone | FXC01452 |
| Description | Stress-induced-phosphoprotein 1 (STI1) (Hsc70/Hsp90-organizing protein) (Hop) (Transformation-sensitive protein IEF SSP 3521) (NY- REN-11 antigen). |
 Length: 1810 bp
Length: 1810 bp Physical map
Physical map
 Restriction map
Restriction map Prediction of protein coding region (GeneMark analysis) for :
cloned DNA seq.
Prediction of protein coding region (GeneMark analysis) for :
cloned DNA seq. Warning for N-terminal truncation: YES
Warning for N-terminal truncation: YES Warning for coding interruption : NO
Warning for coding interruption : NO Integrity of 3' end
Integrity of 3' end
| Length of 3'UTR | 178 bp |
|---|---|
| Genome contig ID | gi51511727f_63617123 |
| PolyA signal sequence (None) |
+----*----+----*----+----*----+---- |
| Flanking genome sequence (111231 - 111280) |
----+----*----+----*----+----*----+----*----+----* |
 Length: 543 aa
Length: 543 aa Result of homology search against nr database
(FASTA output,
Multiple alignment)
Result of homology search against nr database
(FASTA output,
Multiple alignment)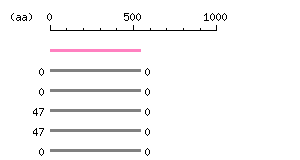 |
|
The numbers on the left and right sides of a black line in the graphical overview indicate the lengths (in amino acid residues) of the non-homologous N-terminal and C-terminal portions flanking the homologous region (indicated by the black line), respectively.
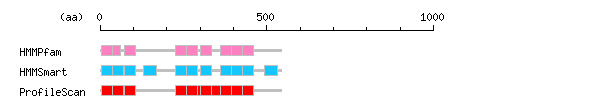
 Result of motif / domain search (InterProScan and SOSUI)
Result of motif / domain search (InterProScan and SOSUI) Result of InterProScan
Result of InterProScan
| Search method | interpro_ID | From | To | Entry | Definition |
|---|---|---|---|---|---|
| HMMPfam | IPR001440 | 4 | 37 | PF00515 | Tetratricopeptide TPR_1 |
| IPR001440 | 38 | 60 | PF00515 | Tetratricopeptide TPR_1 | |
| IPR001440 | 72 | 105 | PF00515 | Tetratricopeptide TPR_1 | |
| IPR013105 | 225 | 258 | PF07719 | Tetratricopeptide TPR_2 | |
| IPR001440 | 259 | 292 | PF00515 | Tetratricopeptide TPR_1 | |
| IPR001440 | 300 | 333 | PF00515 | Tetratricopeptide TPR_1 | |
| IPR001440 | 360 | 393 | PF00515 | Tetratricopeptide TPR_1 | |
| IPR001440 | 394 | 427 | PF00515 | Tetratricopeptide TPR_1 | |
| IPR001440 | 428 | 461 | PF00515 | Tetratricopeptide TPR_1 | |
| HMMSmart | IPR013026 | 4 | 37 | SM00028 | Tetratricopeptide region |
| IPR013026 | 38 | 71 | SM00028 | Tetratricopeptide region | |
| IPR013026 | 72 | 105 | SM00028 | Tetratricopeptide region | |
| IPR006636 | 130 | 169 | SM00727 | Heat shock chaperonin-binding | |
| IPR013026 | 225 | 258 | SM00028 | Tetratricopeptide region | |
| IPR013026 | 259 | 292 | SM00028 | Tetratricopeptide region | |
| IPR013026 | 300 | 333 | SM00028 | Tetratricopeptide region | |
| IPR013026 | 360 | 393 | SM00028 | Tetratricopeptide region | |
| IPR013026 | 394 | 427 | SM00028 | Tetratricopeptide region | |
| IPR013026 | 428 | 461 | SM00028 | Tetratricopeptide region | |
| IPR006636 | 492 | 531 | SM00727 | Heat shock chaperonin-binding | |
| ProfileScan | IPR013026 | 4 | 37 | PS50005 | Tetratricopeptide region |
| IPR013026 | 4 | 105 | PS50293 | Tetratricopeptide region | |
| IPR013026 | 38 | 71 | PS50005 | Tetratricopeptide region | |
| IPR013026 | 72 | 105 | PS50005 | Tetratricopeptide region | |
| IPR013026 | 225 | 258 | PS50005 | Tetratricopeptide region | |
| IPR013026 | 225 | 461 | PS50293 | Tetratricopeptide region | |
| IPR013026 | 259 | 292 | PS50005 | Tetratricopeptide region | |
| IPR013026 | 300 | 333 | PS50005 | Tetratricopeptide region | |
| IPR013026 | 360 | 393 | PS50005 | Tetratricopeptide region | |
| IPR013026 | 394 | 427 | PS50005 | Tetratricopeptide region | |
| IPR013026 | 428 | 461 | PS50005 | Tetratricopeptide region |