Kazusa Clone (Original Type)
Gene/Protein Characteristic Table for
ORK07766
|
Order Kazusa clone(s) from : |
| Accession No | |
|---|---|
| Product ID | ORK07766 |
| Clone name | bm09321 |
| Vector information | |
| Symbol | HEY1 |
| cDNA sequence | DNA sequence (2243 bp) Predicted protein sequence (347 aa) |
|
HaloTag ORF Clone |
FHC07766

|
| Flexi ORF Clone | FXC07766 |
| Description | hairy/enhancer-of-split related with YRPW motif 1 |
 Length: 2243 bp
Length: 2243 bp Physical map
Physical map
 Restriction map
Restriction map Prediction of protein coding region (GeneMark analysis) for :
cloned DNA seq.
Prediction of protein coding region (GeneMark analysis) for :
cloned DNA seq. Warning for N-terminal truncation: NO
Warning for N-terminal truncation: NO Warning for coding interruption : NO
Warning for coding interruption : NO Integrity of 3' end
Integrity of 3' end
| Length of 3'UTR | 1172 bp |
|---|---|
| Genome contig ID | gi51511724r_80738806 |
| PolyA signal sequence (AATAAA,-17) |
+----*----+----*----+----*----+---- |
| Flanking genome sequence (100000 - 99951) |
----+----*----+----*----+----*----+----*----+----* |
 Length: 347 aa
Length: 347 aa Result of homology search against nr database
(FASTA output,
Multiple alignment)
Result of homology search against nr database
(FASTA output,
Multiple alignment)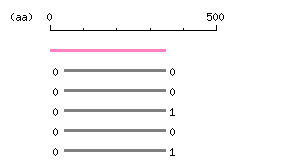 |
|
The numbers on the left and right sides of a black line in the graphical overview indicate the lengths (in amino acid residues) of the non-homologous N-terminal and C-terminal portions flanking the homologous region (indicated by the black line), respectively.
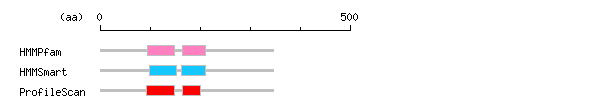
 Result of motif / domain search (InterProScan and SOSUI)
Result of motif / domain search (InterProScan and SOSUI) Result of InterProScan
Result of InterProScan
| Search method | interpro_ID | From | To | Entry | Definition |
|---|---|---|---|---|---|
| HMMPfam | IPR001092 | 95 | 148 | PF00010 | Basic helix-loop-helix dimerisation region bHLH |
| IPR003650 | 164 | 210 | PF07527 | Orange | |
| HMMSmart | IPR001092 | 98 | 153 | SM00353 | Basic helix-loop-helix dimerisation region bHLH |
| IPR003650 | 163 | 210 | SM00511 | Orange | |
| ProfileScan | IPR001092 | 93 | 148 | PS50888 | Basic helix-loop-helix dimerisation region bHLH |
| IPR003650 | 165 | 201 | PS51054 | Orange |